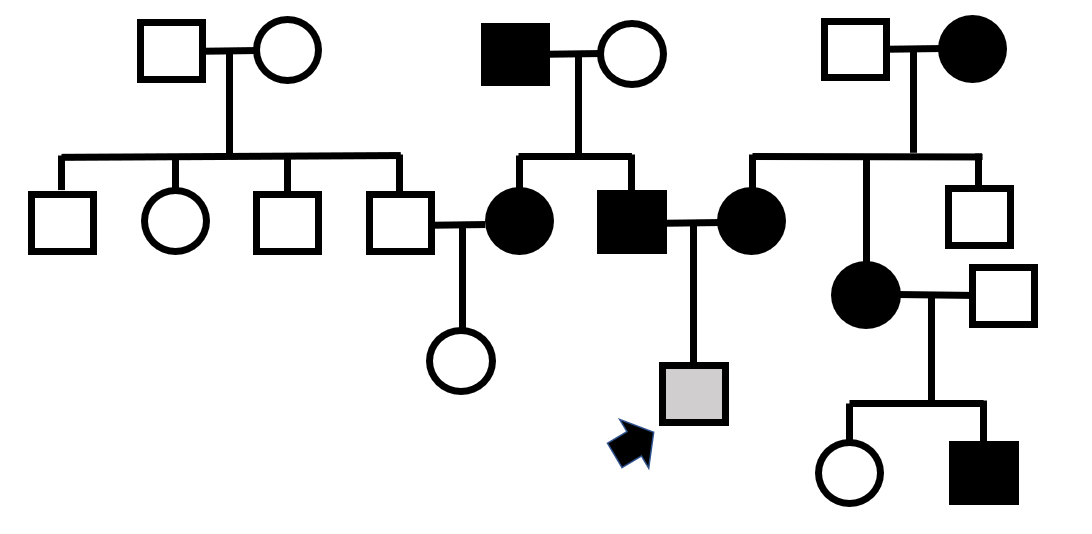
Which mode of inheritance best explains the pedigree above?
Autosomal dominant
X-linked dominant
if it was dominant X-linked then male offspring from affected females would always be affected, the third child of the top right mating is not affected
Non-genetic
there is a genetic explanation for this, try again.
Autosomal recessive
Vaguely possible, but would require a high background carrier rate. There is a more likely explanation.
Mitochondrial
Nope, no suggestion of maternal-only transmission.
What is the the probability that unknown genotype/phenotype person in grey with the arrow will go on to develop the condition?
75%
both parents are heterozygous Aa x Aa. This means you have the possibility of AA, Aa, aA and aa. Three out of four options would develop the disorder and one would be unaffected.
50%
25%
100%
0%
2.5%
really?!
HARD! The affected people get their DNA sequenced and the report identifies the following genes having “variants of unknown significance” (VUS). If you had to predict, based on the inheritance pattern only, which 2 of these changes would be likely to be associated with the disease? Note all are possible, this question is to think about what each of these mutations might mean for protein function.
A single amino acid change in part of a protein involved in polymerisation
We are looking for changes that will exert a dominant effect. Here a change in polymerisation might mean the mutant protein poisons the non mutant protein, i.e. would have dominant effect.
A nonsense (stop codon) mutation in a structural protein
Usually premature termination codon would mean no expression from that allele. Here we are looking at a dominant mutation, most times loss of expression of a structural protein wouldn’t be dominant.
A splice-site mutation affecting a 29 bp exon near the beginning of a gene encoding an enzyme
This might have been tricky, but a splice site mutation often leads to skipping of the affected exon or missplicing. This in turn will lead to a frame-shift if the affected exon is not divisible by 3. The net effect, likely loss of expression. This could be dominant but most likely loss of expression will be recessive unless there is a dosage effect.
A missense mutation affecting the DNA binding site of a transcription factor.
This sort of mutation could be activating; the mutant protein could bind DNA differently / to different sequences and thereby activate the wrong genes or to a different extent. These sort of gain of functions are likely to be dominant.
A truncation mutation to the 3′ untranslated region of a gene involved in extracellular matrix production
often the 3’UTR contains regulatory sequences that define either the stability of the mRNA or possibly where the mRNA is located. These mutations could be dominant but quite often lead to small effects on protein longevity or regulation which might not manifest as a phenotype unless both copies are mutated.
Please note that the third question here was tough when presented as abstract concept. All those mutations could be dominant or recessive depending on context. The key is to consider whether the change will likely cause a gain or loss of function.