This page describes the scientific journey that led to this paper.
Before (and alongside) this study, work on the laminin N terminus (LaNt – friendly intro via link) family of proteins involved observation of where this “new” protein could be found in human tissues, identification that it in breast cancer that levels of the protein are increased (cancer) and also are increased during wound repair. We also did cells-in-a-dish manipulation experiments, where we increased or decreased LaNt expression levels in skin, corneal, or breast cancer cells. Those works indicated that LaNt is capable of influencing cell behaviour including movement and tumour cell invasion.
However, we were missing a big part of the puzzle. The cells in the dish don’t reflect the true complexity of full tissues, while correlation experiments do not imply any causation. So, quite simply, whether or not the LaNt protein is functional in a whole organism was not yet known. This is really important to know; there would be little value in continuing our research in LaNts if they weren’t at least partially functional.
To be able to even begin to answer the questions about whether LaNt is functional in real tissues required a lot of quite complicated work and dedication from two of my PhD students Conor Sugden and. first, Valentina Iorio.
They each constructed what is essentially an artificial gene, then introduced those genes into mice (thereafter the mice are referred to as transgenic).
At this point, I want to acknowledge Prof George Bou Gharios, a world-renowned matrix biologist and genuinely fantastic mentor, friend and colleague. He was a co-supervisor to Valentina and Conor and he and his team brought all their extensive experience to the project.
Episode 1: The Phantom Transgenic
Ethics
Before discussing the how and what of these studies, I want to very briefly address the ethics of using animals. The decision to use animals in research is not trivial. All the experiments have been carried out with a core tenets of “Reduce, Refine, Replace” firmly in the front of our minds. The reason that there were no animal experiments on LaNt proteins up to this point is because we wanted to make sure that the animal studies were required and would be valuable before we begin. We made our in vitro models more and more complex and as accurate as possible by using human material left over from surgery and even eyes from pigs that were being killed for food production (those eyes otherwise go to waste) and got everything we could in vitro. However, only with information from live animals can we be confident that it is worth continuing to study this protein. The UK has some of the tightest laws in the world regarding animal welfare. Everything is planned meticulously to make sure that no animals suffer during the experiments. The studies are designed so that the numbers of animals used are as low as possible yet will provide statistically robust inferences. The animals are housed in custom built specialised facilities with round the clock veterinary staff care. All animals are monitored extensively and no animal is allowed to remain in pain. Before any work could start, extensive study plans were independently assessed by the home office, edited in light of their recommendations and throughout the studies we reported all events. The ethics approval process took over 6 months, which is pretty standard in the UK.
The best laid plans…
The original plan (c2016) was to express the transgene only in epithelial cells as our work at that point had focused on the skin skin and front of the eye epithelium. To achieve expression only in these places, we put our LaNt transgene under the control of the genetic elements that regulate expression of a skin keratin gene. Valentina made this construct and then extensively tested it in cells and a dish and everything worked perfectly. However, it turned out that we couldn’t generate animals expressing this construct.
Our very strong feeling was that expression of the transgene was preventing normal development. However, without the animals to analyse, we couldn’t know what the problem was. You can’t publish phantom transgenics!
For Valentina, this was the end of her PhD funding. She also did a lot of the cell culture work and was able to produce a really lovely doctoral thesis and flew through her viva in 2017.
So, what to do next? Conor joined the team in October 2017 and we started again (of course).
A little side note, perseverance is one of those traits that you look for in employees. Most of scientific research is a battle between your desire to get the answer versus your frustration at how difficult or slow things are going. The question behind this research was important to answer and worth pursuing. In both Valentina and Conor I have been lucky to have two people who were willing to keep on going even when things were really frustrating!
Episode II: Attack of the Clones*
*The process of making these constructs is known as cloning. It can be quite slow, especially when you need to insert multiple different things into the same construct.
The new plan involved making a new gene construct with some extra control features. We switched the promoter, the driver of the gene expression, to one that is active in every cell. However, really importantly, we slotted in a “STOP” signal between the promoter and the LaNt gene. This stop signal meant that although the mice would have the transgene it wouldn’t actually make any protein until the stop cassette was removed. This addition gave us more control over expression.
To remove the stop cassette in a controlled manner we use a second transgene to express an enzyme (Cre recombinase) that can chop out the bit of genetic code containing the stop. Once that happens a cell will start making the LaNt protein from that point onward.
This is a really flexible system; it allows us to selectively turn on LaNt expression in different cell types depending on where and when the other transgene is expressed. In the case of the published study, our second transgene was widely expressed but the enzyme only becomes active once tamoxifen is added. Therefore this system allowed for temporal control of when expression started as we could choose when to inject the animals with tamoxifen.
We also built in some extra features in the LaNt construct to allow us to know which cells were expressing the protein. A little bit of fancy genetics allows two proteins to be produced at once but are made separate from each other. We used this feature to allow the transgene to produce the LaNt protein and also a fluorescent red protein called tdTomato. With this addition, we would know that any cell that was red was also making LaNt.
Episode III: Revenge of the blot
Before even thinking about moving into animals, Conor conducted a whole battery of in vitro tests. Here, as the construct is quite complicated with potential for not working exactly as planned for a variety of reasons, the testing was quite extensive. In one of the tests, shown below, he checked that the new LaNt construct was expressed in cells only after exposure to the Cre enzyme to remove the stop cassette. To do so, he set up the four conditions in a epithelial cell line, using a green protein tagged Cre (GFP) to visualise cells expressing Cre and using the tdTomato as a read-out for the LaNt protein. In the pictures you can see that the only time the tdTomato signal is observed is in the bottom row, where the cells have both the Cre and the LaNt construct. The little picture to the bottom right shows you a close up of on cell, with the Cre in the nucleus in green and the rest of the cell filled up with TdTomato (in purple). This experiment + some others that are less visually interesting combined to demonstrate that the constructs worked as we designed them to.


The “some others” experiment included “western blots”, a method to detect protein using antibodies. I think Conor would freely admit that he isn’t a fan of “blotting”, it took him quite awhile to get the data we needed to be convinced things were ready to move on.
Episode IV: A New Hope
Once we were 100% convinced that everything was working, we progressed to introducing the new construct into mouse eggs and implanting them into recipient mothers. Basically, mouse IVF. Once the mice gave birth, Conor was able to test whether the transgene was inserted or not using PCR and sequencing. Then he tested whether the introduced gene still “worked” (i.e., made the protein ) by isolating cells from the animals and treating them with the Cre enzyme. The big difference in this second round of tests was that these cells actually came from mice where the transgene was part of their DNA, i.e. would be passed on when mated, rather than being introduced temporarily.
And good news. It worked! About a year of Conor’s PhD was over but we had animals that were now ready for the next stage.
Next up was to mate our LaNt transgenic animals with another mouse line that could express the Cre enzyme. There are lots of options for this, this Cre/Lox system has been widely used and there lots of lines available where the Cre expressed in different tissues or at different times. The one we selected is known as Rosa26-CreERT. This one expresses everywhere but only once tamoxifen has been injected. The tamoxifen activates the enzyme by allowing it to move from the cell membrane into the nucleus where it can act and cut out the STOP cassette.
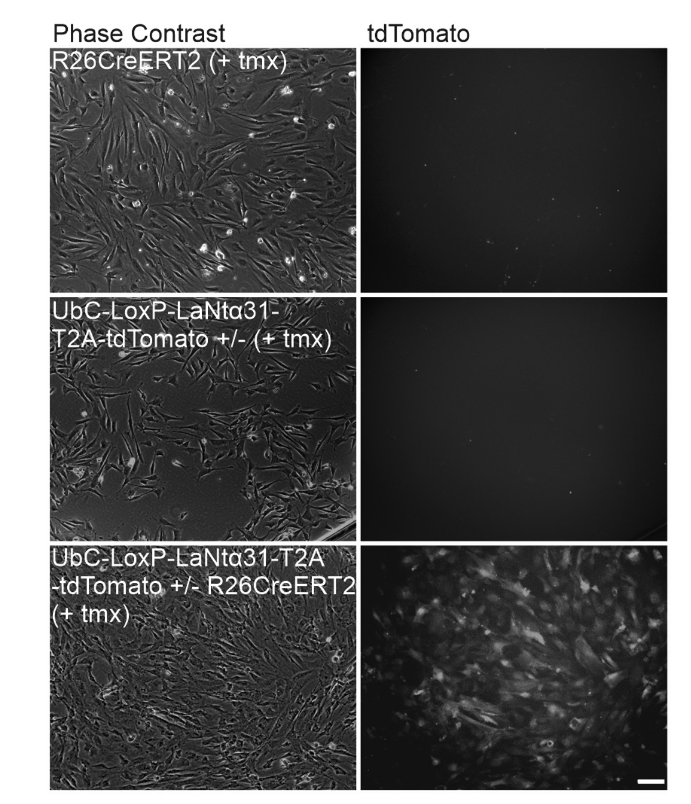
After crossing the two lines we were able to test if the cells from double transgenic animals (now LaNt + Cre) did what we hoped. The images below are from those animals. Top row is the Cre only line, middle is the LaNt only line and bottom is the LaNt+Cre line. In each case, tamoxifen has been added. As you can see, the tdTomato as a marker for the LaNt protein, is only present in the double transgenics.
For info- a mouse gestation period is 3 weeks, and they take 6 weeks from birth to becoming an adult. Part of the reason this sort of work takes so long is that you are waiting for breeding cycles, for animals of the correct genotype in the correct ratios. It is all very carefully controlled and we are at the mercy of the biology and statistics for what we get and when.
Fantastic news! We had double transgenics that express the LaNt protein when exposed to tamoxifen. Now all we have to do is decide on the experimental set up in terms of timing and dosing to determine what happens.
On we go.
Or did we?
Episode V: COVID strikes back
We’d actually just started the functional studies in early 2020 when, as you know, the COVID pandemic struck. Our University shut down for 3 months and the animal house decided it was ethically best to reduce all animal colonies during that time as we didn’t how long things would be affected. All experiments stopped.
When the labs opened up again, we needed to expand the colony again before we had sufficient animals to do our studies. Those 6 weeks waiting for animals to age were adding up!
Episode VI: Return of the Transgenic
Finally. Mid 2020 and we were able to find the answer to the question “what happens if you increase LaNt expression during development?”. We treated pregnant mothers with tamoxifen at embryonic day 14 and then waited for them to give birth 7 days later.
The results were striking. Immediately at birth some of the animals had visible red regions in their skin, areas where blood vessels had leaked. We checked the animals and those leaky blood vessels only ever happened in the mice that expressed the LaNt protein.
This is actually a huge result. It says in plain and simple terms that too much LaNt protein disrupts blood vessel function.
What followed was a whole load of further tests to see exactly how the bad the situation was, which tissues were affected and when. Some images are below.
Really striking was the kidneys. When Conor dissected out the kidneys, there was massive colour difference between the LaNt transgenics and the normal animals, and when we looked closer at the histology it was apparent that there was lots of blood outside of vessels in the kidney (yellow arrows point to bright pink blood cells in the tissue stain).

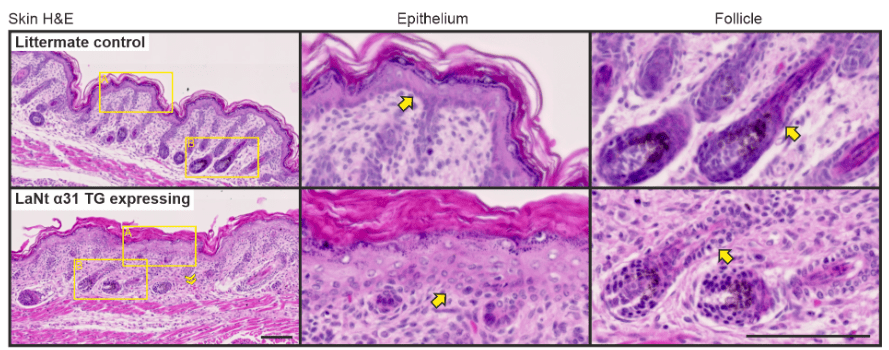
In the skin the disruption was less dramatic. There were areas with blood cells visible but mostly the tissue looked OK at first glance. Compare top and bottom of the pics below. Right is a hair follicle. Most noticeable is the bit where the yellow arrows are pointing, the junction between the cell sheets of the epithelium and the underlying tissue. This junction “the dermal-epidermal junction” is the skin epithelial basement membrane.
We did a few other experiments to look more closely at that junction. These included transmission electron microscopy. What you can see below is zoomed in really close to that junction. A small part of the bottom of one cell runs along the top with the matrix beneath. The red chevrons point to structures known as hemidesmosomes. These are kind of like spot-weld points that button down the epithelium onto the matrix. Here we saw that the hemidesmosomes were larger and more mature individually but also that there were slightly fewer. We’ve done a bunch of cell culture experiments that show the same thing, keep your eyes out for that paper later in 2022 (hopefully).

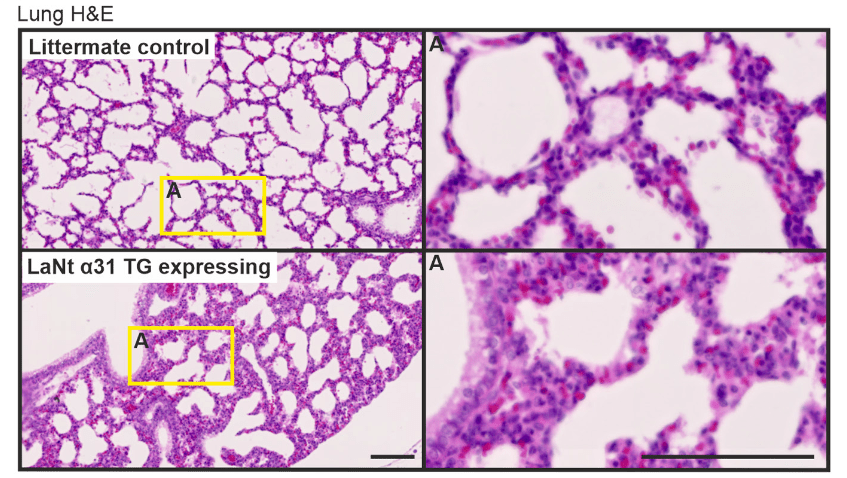
Conor also looked at lungs and liver. The most interesting thing here was that the number of cells in the liver were reduced in the LaNt overexpressing animals. This likely indicates an effect on the local stem cell niche or stem cell population and isn’t something we really anticipated, we are excited to follow up on this discovery.

Episode VII: Reviewers awaken
We actually had almost all these results very soon after the labs opened up again. We submitted the manuscript in December 2020 and awaited the verdict of our peers.
Peer review usually makes the paper better in the long run and this was definitely true here. However, it took forever. In the first round of review some additional experiments were requested including the electron microscopy studies. This actually entailed new animals being generated and analysed and took 6+ months before we could resubmit. After resubmission, the reviewers looked at the updated manuscript and still weren’t quite happy, so more experiments were performed and the text updated. And then the same cycle happened again. Eventually, after much sweat and tears (and extravascular blood!), the manuscript was accepted in April 2022. 6 years after we began.
The wait still wasn’t over though, not really clear why, but there was another 2 month delay from acceptance to publication on June 1st 2022.
Conor also submitted his thesis on April 30th and has his viva at the end of the month prior to starting a postdoctoral position in Barcelona.
The saga continues
It has been a saga but now that this paper is out, the doors are open for the follow up work (Episode VIII, IX, + spin offs). There is a lot more to do now that the findings from these studies tell us that this protein, that no one had ever studied in live animals before, is really potent. It changes the way we think about how cell-matrix interactions are regulated and has generated more questions for us and others to pursue.
Footnotes
Some of the work my lab does is focused on developing new treatments, it is objective-driven research and the motivation is really strong, each step takes you closer to improving the lives of people both now and in the future. Other projects, like this one, are discovery science; the true value won’t be known until a long time in the future. All the objective-driven research and treatments being developed today are built upon discoveries made many years ago. You can’t get translation without discovery. Slightly different mindsets are required for the different projects; for translational work you are looking for “the solution ” to a problem, for discovery you are looking for “the answer” to a question.