This page is aimed at a non-specialist audience, summarising recent research into breast cancer and LaNt a31. The primary-data peer-reviewed paper can be accessed via this link (free).
Breast cancer is extremely common, so much so that I can confidently assert that anyone reading this blog will know someone has been affected. However, when we talk about breast cancer we aren’t really talking about one uniform disease; it varies dramatically between individuals. Those individual differences are not only how much the disease has progressed, but also “molecular drivers”, the specific set of proteins or genetic differences that are responsible for the disease. Knowing what is driving the disease can help medical professionals decide on a personalised course of treatment; choosing interventions to target the right pathways, or even choosing between different intensities of the same treatment dependent on how aggressive the disease is likely to be. Collectively we refer to these as biomarkers.
There are lots of valuable cancer biomarkers already available. When you hear about “triple negative” breast cancer, that refers to three biomarkers that have been used for a long time already and some really powerful drugs have been developed to target these markers. However, identification of additional, new biomarkers can help to further stratify patients and improve treatment decisions. Moreover, identifying whether a protein that contributes to the progression of the disease, can mean that we have a new target for therapeutic intervention.
With these points in mind, we set out investigating a protein that no one had ever studied before in breast cancer, LaNt a31.
Laminin N terminus alpha31 (LaNt a31)
We’ve written quite a lot of background on LaNt a31 in other places on this blog, here are some of the important and relevant points from those studies with links to the non-specialist descriptions
- LaNt a31 is the newest* member of the laminin superfamily- *in terms of evolution and discovery.
- The laminins have been extensively studied: they are essential extracellular matrix / basement membrane proteins with important roles in many processes including cell adhesion, cell migration, as barriers to tumour invasion and in regulating cellular proliferation and differentiation. Dysregulation of laminin expression is a feature of most solid cancer subtypes and they have been investigated for biomarker potential in lots of studies.
- LaNt a31 in contrast has been studied a lot less!
- LaNt a31 is derived by a rare form of alternative splicing (intron retention and alternative polyadenylation) from the LAMA3 gene.
- LaNt a31 is widely expressed in normal human tissues including around terminal ductal lobular units in the breast.
- LaNt a31 is upregulated during corneal burn wound repair and regulates epithelial cell migration in cells in a dish.
- We hypothesise that LaNt a31 can change laminin organisation, which in turn changes the ways cells respond to laminins.
LaNt a31 expression is increased in breast cancer
Like most studies, our work on LaNt a31 began with a pilot study. We obtained a small panel of uninvolved (normal) and breast cancer specimens that have been donated to our local research tissue bank. We then used antibodies that we raised and validated against LaNt a31 to localise where and how much of the protein was present in each tissue using a process of “immunohistochemistry” (literally using antibodies (immuno) to probe tissue (histo) and detecting them with a chemical reaction).
In the images below, the brown stain comes from an enzymatic reaction where the antibodies are bound to their target. Broadly speaking, the more brown, the more antibodies are bound and, therefore, the more protein that is present. This type of quantification is not particularly precise in terms of absolute abundance but it does give a reasonable indication of relative abundance. In this small scale study, all the normal tissue specimens gave very little staining in very discrete regions, whereas most of the cancer specimens were strongly stained (examples below). These findings suggested that LaNt a31 might be upregulated in breast cancer and that it would be worth looking at a larger cohort…. so we did!
To increase the sample size, we turned to tumour microarrays. These contain a small pieces from 50-200 specimens all on a single slide. The big advantage of this approach is that the processing steps are standardised across all the specimens and this makes it fairer to compare staining intensity as an indicator of expression level. The disadvantage of this array approach is that you examine only a very small part of a tumour and could miss the interesting bit (experimental design is all about compromises). However, the pilot data had suggested that the increase in LaNt a31 expression wasn’t restricted to a specific part of the tumours. Therefore, the array approach looked promising to answer the next broad question: is LaNt a31 upregulated in breast cancer?
To answer this first question, we processed arrays containing matched samples from a tumour and from normal tissue that came from the same person. This allowed us to directly compare the staining intensity between the paired samples and from those scored determine whether the expression was either increased, decreased or unchanged in tumour.
Throughout these analyses we used a “blinded” scoring system where the staining was done by one researcher (Lee Troughton) and the intensity scored by two or more other people who didn’t have any prior information about the samples. The clinical features of each specimen where only revealed after the staining scores were assigned. We used multiple scorers to reduce the impact of subjectivity upon the scoring. We designed in these approaches reduce possibility of bias and thereby increase confidence in the results obtained.
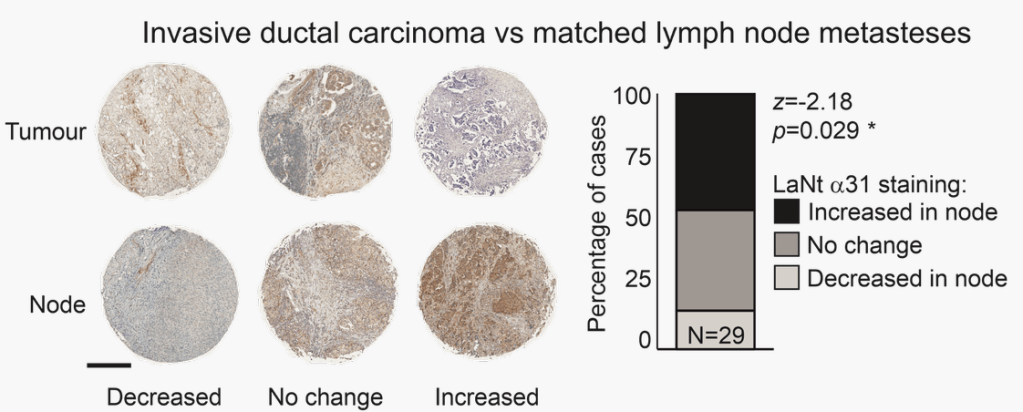
Analysis of these matched normal vs tumour data revealed that in more than half the breast cancer samples analysed, the LaNt a31 staining was darker in the tumour than normal tissue.
We then compared matched primary tumour vs the distal nodal metastasis in the same person. Again, these analysis identified higher LaNt a31 expression in the sample from the metastasis compared with the paired primary tumour.
From these studies, we conclude that LaNt a31 is upregulated in breast cancer and that LaNt a31 is upregulated in breast cancer metastases compared with primary tumour.

LaNt a31 as breast cancer biomarker
We then processed a much larger set of patient samples where we had information about the outcomes of the patients and lots of other clinical data.
This analysis was actually a little disappointing (and surprising). It revealed that there actually wasn’t a strong association between tumour grade (how “bad” a tumour looks) and LaNt a31 staining intensity (B below) or LaNt expression and the presence of nodal involvement (D below). However, we did see that the highly proliferative tumours were likely to have high LaNt a31 staining (C below, Ki67 is a proliferation marker).
When we looked at patient survival, we observed a slight difference between the highest and lowest expression. However, this observation a p value of 0.13*. Therefore, although we have observed a difference we are not very confident about this as result

The whole population data that we got from these sample is interesting; however, we cannot forget that each breast cancer is different from the next. Therefore, we also interrogated the data for any correlation between the expression of known drivers of breast cancer and LaNt a31 expression levels. These additional analyses revealed a positive association between epidermal growth factor receptor and LaNt a31 and a negative association between estrogen receptor expression and LaNt a31.
More exciting; in one third of the “triple negative” breast cancers cases there was very high LaNt a31 expression. “Triple negative” refers to absence of expression of three of the main therapeutic targets for breast cancer. This subset of tumours are often more difficult to treat and there is a pressing need to identify new drug targets. The increased LaNt a31 expression, even within only a subset of these patients, might be something we can exploit in the future as a target for new drugs.
But what does it mean when LaNt a31 expression increases
As anyone who’s ever taken a research design class will recite, “correlation does not imply causation”. They’ll also tell that correlation does not tell you about directionality and they might mention something about Nicolas Cage movies and swimming pool deaths (aka spurious correlations).
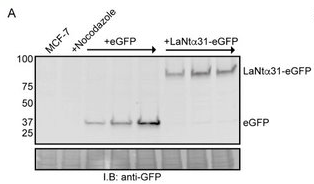
Therefore our next series of experiments were designed to directly investigate the effect of increased LaNt a31 expression upon breast cancer. To do this, we made an artificial construct (similar to how the RNA vaccines work) to allow us to induce LaNt a31 expression in breast cancer cells in a dish. In our construct, we tagged the LaNt a31 with a green fluorescent protein (GFP) on one end, this allowed us to track which cells were expressing the construct and also how high the expression was. We also could then use cells expressing a construct for just the GFP protein as a “control” or comparison treatment.
Using these tools, we were asked whether increasing LaNt a31 expression directly affects core cell behaviours associated with breast cancer .
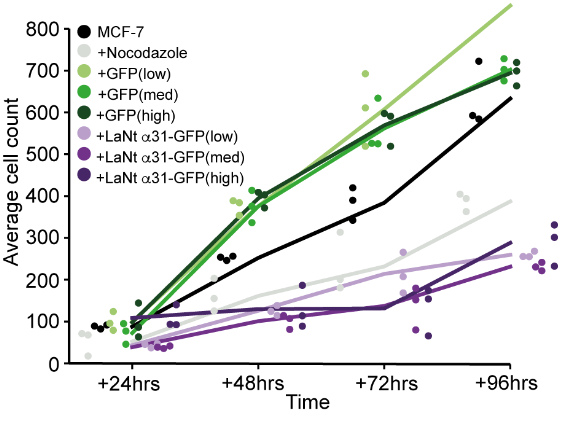
First we asked what effect does increased LaNt a31 expression have upon cell proliferation. We observed that the cells expressing the LaNt construct (purple) proliferated at a reduced rate compared with the control treated cells (green, black). This was a little surprising – our tissue staining suggested that it would be the opposite way around (don’t you just love science!). We also tried some extra experiments in an attempt to work out if expression goes the other way around i.e. if high proliferation drives higher LaNt expression. Those data said proliferation is not required but the jury is out whether its a driver… watch this space.
We next asked whether induced LaNt a31 expression influenced the way in which the cancer cells moved. We did these experiments in multiple different ways; investigating sheet-like migration, single cell migration and invasion into different matrixes. We performed these experiments with two very different cell lines; MCF7 which are low movement and non-invasive line that model a not very aggressive cancer and MDA-MB-231 which are the opposite, derived from a highly invasive cancer! Each of the different types assays combined with the different cell types tells us something slightly different about how LaNt a31 expression affects the cells, and then the individual experiments combine to provide a more complete picture than performing any one experiment on its own.

Loads of experiments later (this post is already getting long) and we conclude that … LaNt a31 expression doesn’t have a major effect on speed of migration (perhaps slightly slower).
BUT
LaNt a31 changed the way the cells invaded.
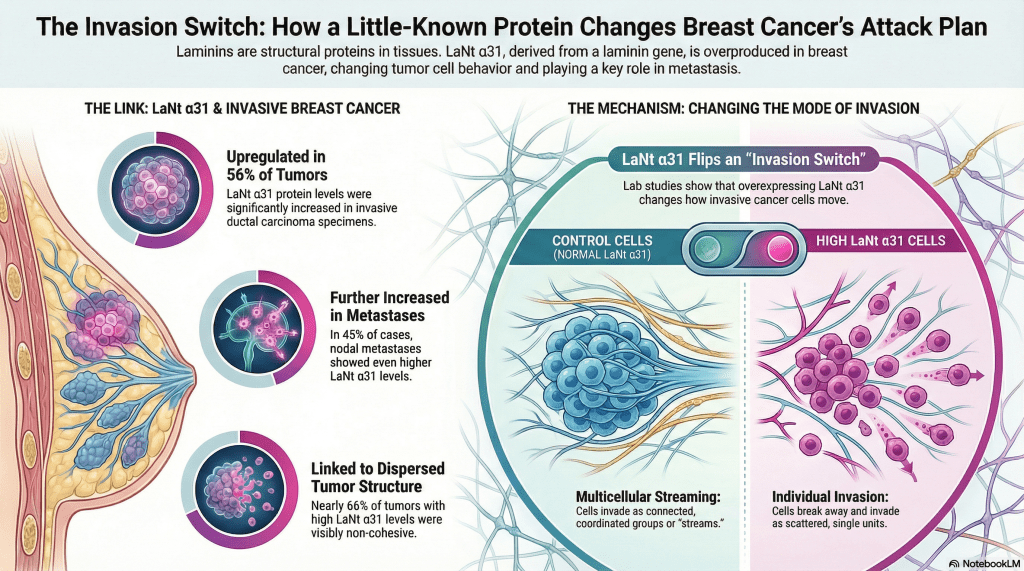
Normally, MDA-MB-231 invasive cells invade as strings of cells, follow my leader style “multicellular streaming”. In the LaNt a31 expressing cells this changed to a different form of invasion, with each individual cell apparently acquiring the ability to invade. Extremely interesting! But there’s more. That change in invasion “style” was only observed when the cells were migrating into a laminin containing matrix and not when the identical assay was performed when migrating into collagen. Remember how I said earlier that LaNt a31 is laminin-related and we hypothesised it would work on laminins…


These findings of an apparently laminin-dependent effect of LaNt a31 upon tumour invasion is something that we are working on now (Fawziah Asiri’s PhD project) and what it means in terms of how breast cancers develop or progress is something that we really need to understand.
In the short-term, this finding of change in invasion style led us to reassess some of the tumours we had analysed earlier. Specifically, we looked at those tumours with high LaNt expression and we observed that they looked different, less cohesive which implies that there these in vitro observations could also be reflected in real-tissue behaviour.
We could do more, indeed we are doing more now, but we decided that these findings are interesting enough that it was worth sharing in their current form.
Read the paper
You can read the published work here (takes you to PLOSone – open access)
You can also read about some of our other work on LaNts here (cancer), here (distribution), here (corneal wound healing) and here (general intro). Or check the blog for other work being done by the team.
*P values indicate type I error – essentially the false positive rate. The lower the p value, the more confident you can be that the observation a true reflection of the wider population. People often use thresholds such as p below 0.05 (5%) or below 0.01 as cut-offs for what sort of result should be trusted.